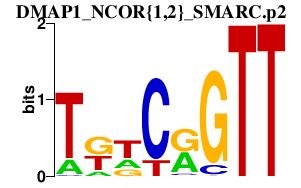
|
chr1_-_157772420
|
0.902
|
NM_001004469
|
OR10J5
|
olfactory receptor, family 10, subfamily J, member 5
|
|
chr3_+_120799411
|
0.820
|
NM_015900
|
PLA1A
|
phospholipase A1 member A
|
|
chr8_-_114518194
|
0.820
|
NM_052900
NM_198123
|
CSMD3
|
CUB and Sushi multiple domains 3
|
|
chr1_-_47468029
|
0.619
|
NM_003189
|
TAL1
|
T-cell acute lymphocytic leukemia 1
|
|
chr9_-_8723893
|
0.606
|
NM_001040712
NM_001171025
NM_130391
NM_130392
NM_130393
|
PTPRD
|
protein tyrosine phosphatase, receptor type, D
|
|
chr12_+_99712504
|
0.568
|
NM_178826
|
ANO4
|
anoctamin 4
|
|
chr5_+_32824701
|
0.537
|
NM_024563
|
NPR3
|
natriuretic peptide receptor C/guanylate cyclase C (atrionatriuretic peptide receptor C)
|
|
chr6_-_48186868
|
0.493
|
|
C6orf138
|
chromosome 6 open reading frame 138
|
|
chr18_+_59374372
|
0.491
|
NM_080474
|
SERPINB12
|
serpin peptidase inhibitor, clade B (ovalbumin), member 12
|
|
chr13_-_46368968
|
0.473
|
NM_000621
NM_001165947
|
HTR2A
|
5-hydroxytryptamine (serotonin) receptor 2A
|
|
chr1_-_93030538
|
0.471
|
NM_005665
|
EVI5
|
ecotropic viral integration site 5
|
|
chr4_+_93444572
|
0.470
|
NM_001510
|
GRID2
|
glutamate receptor, ionotropic, delta 2
|
|
chr7_-_30032649
|
0.464
|
NM_017946
|
FKBP14
|
FK506 binding protein 14, 22 kDa
|
|
chr5_+_156540425
|
0.462
|
NM_005546
|
ITK
|
IL2-inducible T-cell kinase
|
|
chrX_+_78087484
|
0.456
|
NM_014499
NM_198333
|
P2RY10
|
purinergic receptor P2Y, G-protein coupled, 10
|
|
chr11_-_104477367
|
0.438
|
NM_001007232
|
CARD17
|
caspase recruitment domain family, member 17
|
|
chr17_+_35038276
|
0.422
|
NM_181505
|
PPP1R1B
|
protein phosphatase 1, regulatory (inhibitor) subunit 1B
|
|
chr17_+_55854636
|
0.417
|
NM_181707
|
C17orf64
|
chromosome 17 open reading frame 64
|
|
chr10_+_18280773
|
0.413
|
NM_001145195
NM_152725
|
SLC39A12
|
solute carrier family 39 (zinc transporter), member 12
|
|
chr11_-_5302157
|
0.405
|
NM_033180
|
OR51B2
|
olfactory receptor, family 51, subfamily B, member 2
|
|
chr3_+_182900081
|
0.403
|
|
SOX2OT
|
SOX2 overlapping transcript (non-protein coding)
|
|
chr6_+_52159143
|
0.403
|
NM_002190
|
IL17A
|
interleukin 17A
|
|
chr6_-_33016561
|
0.396
|
NM_002118
|
HLA-DMB
|
major histocompatibility complex, class II, DM beta
|
|
chr4_-_116254381
|
0.390
|
NM_022569
|
NDST4
|
N-deacetylase/N-sulfotransferase (heparan glucosaminyl) 4
|
|
chr4_+_134289893
|
0.382
|
NM_020815
NM_032961
|
PCDH10
|
protocadherin 10
|
|
chr17_-_31231421
|
0.381
|
NM_002985
|
CCL5
|
chemokine (C-C motif) ligand 5
|
|
chr4_-_153820535
|
0.379
|
NM_152680
|
TMEM154
|
transmembrane protein 154
|
|
chr3_-_42789738
|
0.375
|
NM_144719
|
CCDC13
|
coiled-coil domain containing 13
|
|
chr1_+_57093030
|
0.371
|
NM_000562
|
C8A
|
complement component 8, alpha polypeptide
|
|
chr5_-_145232723
|
0.370
|
NM_001080516
|
GRXCR2
|
glutaredoxin, cysteine rich 2
|
|
chr7_+_101978777
|
0.367
|
NM_001166339
NM_001031618
|
SPDYE2L
SPDYE2
|
WBSCR19-like protein 3
speedy homolog E2 (Xenopus laevis)
|
|
chr1_+_100957883
|
0.364
|
NM_001078
NM_080682
|
VCAM1
|
vascular cell adhesion molecule 1
|
|
chr14_+_36196523
|
0.356
|
NM_006194
|
PAX9
|
paired box 9
|
|
chr11_+_130745580
|
0.356
|
NM_001048209
|
NTM
|
neurotrimin
|
|
chr15_+_42815851
|
0.356
|
NM_080745
NM_182985
|
TRIM69
|
tripartite motif containing 69
|
|
chr3_+_125586241
|
0.354
|
|
KALRN
|
kalirin, RhoGEF kinase
|
|
chr1_-_57204208
|
0.350
|
NM_000066
|
C8B
|
complement component 8, beta polypeptide
|
|
chr16_-_51139214
|
0.350
|
NM_001146188
|
TOX3
|
TOX high mobility group box family member 3
|
|
chr4_+_14950657
|
0.345
|
NM_001135170
|
C1QTNF7
|
C1q and tumor necrosis factor related protein 7
|
|
chr4_+_147316284
|
0.344
|
NM_007080
|
LSM6
|
LSM6 homolog, U6 small nuclear RNA associated (S. cerevisiae)
|
|
chr16_-_4778317
|
0.343
|
NM_001154458
NM_144605
|
SEPT12
|
septin 12
|
|
chr5_-_27074407
|
0.343
|
NM_016279
|
CDH9
|
cadherin 9, type 2 (T1-cadherin)
|
|
chr22_+_38652555
|
0.338
|
|
GRAP2
|
GRB2-related adaptor protein 2
|
|
chr1_-_198643885
|
0.337
|
|
ZNF281
|
zinc finger protein 281
|
|
chr13_+_23042722
|
0.335
|
NM_148957
|
TNFRSF19
|
tumor necrosis factor receptor superfamily, member 19
|
|
chr22_-_27787793
|
0.334
|
NM_015370
|
C22orf31
|
chromosome 22 open reading frame 31
|
|
chr12_+_381001
|
0.333
|
NM_001130146
NM_032358
|
CCDC77
|
coiled-coil domain containing 77
|
|
chr5_+_133870175
|
0.331
|
|
|
|
|
chr2_+_65517316
|
0.329
|
|
FLJ16124
|
FLJ16124 protein
|
|
chr11_+_63060848
|
0.328
|
NM_004585
|
RARRES3
|
retinoic acid receptor responder (tazarotene induced) 3
|
|
chr4_-_169476391
|
0.325
|
NM_017631
|
DDX60
|
DEAD (Asp-Glu-Ala-Asp) box polypeptide 60
|
|
chr17_-_78000890
|
0.315
|
NM_001193655
|
C17orf62
|
chromosome 17 open reading frame 62
|
|
chrX_+_119379967
|
0.314
|
NM_001142447
NM_012069
|
ATP1B4
|
ATPase, Na+/K+ transporting, beta 4 polypeptide
|
|
chr4_-_174774368
|
0.312
|
NM_006792
|
MORF4
|
mortality factor 4
|
|
chr3_-_188903038
|
0.307
|
NM_001004312
|
RTP2
|
receptor (chemosensory) transporter protein 2
|
|
chr2_-_200033445
|
0.307
|
|
SATB2
|
SATB homeobox 2
|
|
chr11_-_84312085
|
0.306
|
NM_001364
|
DLG2
|
discs, large homolog 2 (Drosophila)
|
|
chr14_+_75907442
|
0.306
|
NM_004452
|
ESRRB
|
estrogen-related receptor beta
|
|
chr14_-_69953433
|
0.305
|
NM_018373
|
SYNJ2BP
|
synaptojanin 2 binding protein
|
|
chr10_-_69125932
|
0.303
|
NM_013266
|
CTNNA3
|
catenin (cadherin-associated protein), alpha 3
|
|
chr13_-_25687062
|
0.302
|
|
RNF6
|
ring finger protein (C3H2C3 type) 6
|
|
chr6_-_52217185
|
0.302
|
NM_052872
|
IL17F
|
interleukin 17F
|
|
chr12_+_94391952
|
0.296
|
NM_006838
|
METAP2
|
methionyl aminopeptidase 2
|
|
chr9_+_106371269
|
0.295
|
NM_001004483
|
OR13C8
|
olfactory receptor, family 13, subfamily C, member 8
|
|
chr17_-_30414789
|
0.295
|
NM_001017368
|
RFFL
|
ring finger and FYVE-like domain containing 1
|
|
chr1_-_171838802
|
0.294
|
NM_178527
|
SLC9A11
|
solute carrier family 9, member 11
|
|
chr7_+_139750311
|
0.292
|
NM_001008749
|
RAB19
|
RAB19, member RAS oncogene family
|
|
chr6_-_46811388
|
0.291
|
NM_001168357
|
PLA2G7
|
phospholipase A2, group VII (platelet-activating factor acetylhydrolase, plasma)
|
|
chr8_+_11877238
|
0.291
|
NM_001033017
|
DEFB135
|
defensin, beta 135
|
|
chr12_+_106692291
|
0.290
|
NM_203436
|
ASCL4
|
achaete-scute complex homolog 4 (Drosophila)
|
|
chr3_+_123279439
|
0.289
|
NM_006889
|
CD86
|
CD86 molecule
|
|
chr21_-_30614477
|
0.287
|
NM_203405
|
KRTAP26-1
|
keratin associated protein 26-1
|
|
chr2_+_61986287
|
0.286
|
NM_152516
|
COMMD1
|
copper metabolism (Murr1) domain containing 1
|
|
chr5_-_22248630
|
0.285
|
|
CDH12
|
cadherin 12, type 2 (N-cadherin 2)
|
|
chr14_-_34252695
|
0.281
|
NM_021914
|
CFL2
|
cofilin 2 (muscle)
|
|
chr1_-_103346482
|
0.281
|
|
COL11A1
|
collagen, type XI, alpha 1
|
|
chr4_+_3220564
|
0.280
|
NM_001042690
|
C4orf44
|
chromosome 4 open reading frame 44
|
|
chr2_-_37752731
|
0.279
|
NM_006449
|
CDC42EP3
|
CDC42 effector protein (Rho GTPase binding) 3
|
|
chr4_+_88187215
|
0.279
|
|
AFF1
|
AF4/FMR2 family, member 1
|
|
chr4_-_145281237
|
0.278
|
NM_002099
|
GYPA
|
glycophorin A (MNS blood group)
|
|
chr16_+_24475129
|
0.275
|
|
RBBP6
|
retinoblastoma binding protein 6
|
|
chr17_+_22823151
|
0.275
|
NM_014238
|
KSR1
|
kinase suppressor of ras 1
|
|
chr11_-_5321329
|
0.271
|
NM_001005567
|
OR51B5
|
olfactory receptor, family 51, subfamily B, member 5
|
|
chr18_-_68683913
|
0.270
|
NM_138999
NM_153181
|
NETO1
|
neuropilin (NRP) and tolloid (TLL)-like 1
|
|
chr7_-_27186400
|
0.269
|
NM_153715
|
HOXA10
|
homeobox A10
|
|
chr4_+_129021514
|
0.267
|
NM_001190801
|
PLK4
|
polo-like kinase 4
|
|
chr2_-_74424041
|
0.267
|
NM_133478
|
SLC4A5
|
solute carrier family 4, sodium bicarbonate cotransporter, member 5
|
|
chr11_+_120478584
|
0.266
|
NM_005422
|
TECTA
|
tectorin alpha
|
|
chr18_-_41923571
|
0.263
|
|
ATP5A1
|
ATP synthase, H+ transporting, mitochondrial F1 complex, alpha subunit 1, cardiac muscle
|
|
chr8_+_142501188
|
0.263
|
NM_007079
NM_032611
|
PTP4A3
|
protein tyrosine phosphatase type IVA, member 3
|
|
chr17_-_46140015
|
0.262
|
NM_052855
|
ANKRD40
|
ankyrin repeat domain 40
|
|
chr11_-_84311867
|
0.260
|
|
DLG2
|
discs, large homolog 2 (Drosophila)
|
|
chr1_-_103346639
|
0.260
|
NM_001190709
NM_001854
NM_080629
NM_080630
|
COL11A1
|
collagen, type XI, alpha 1
|
|
chr17_-_44642934
|
0.256
|
NM_001198754
|
GNGT2
|
guanine nucleotide binding protein (G protein), gamma transducing activity polypeptide 2
|
|
chr9_-_32516164
|
0.255
|
NM_014314
|
DDX58
|
DEAD (Asp-Glu-Ala-Asp) box polypeptide 58
|
|
chr1_+_178190493
|
0.252
|
NM_014810
|
CEP350
|
centrosomal protein 350kDa
|
|
chr19_+_17377502
|
0.249
|
|
LOC100507535
|
hypothetical LOC100507535
|
|
chr7_-_113346284
|
0.245
|
NM_002711
|
PPP1R3A
|
protein phosphatase 1, regulatory (inhibitor) subunit 3A
|
|
chr4_+_39874921
|
0.244
|
NM_004310
|
RHOH
|
ras homolog gene family, member H
|
|
chr2_-_215711313
|
0.244
|
NM_173076
|
ABCA12
|
ATP-binding cassette, sub-family A (ABC1), member 12
|
|
chr11_-_88864061
|
0.243
|
NM_001143836
NM_016931
|
NOX4
|
NADPH oxidase 4
|
|
chr2_-_68952006
|
0.243
|
NM_014482
|
BMP10
|
bone morphogenetic protein 10
|
|
chr4_+_114190233
|
0.243
|
NM_001148
NM_020977
|
ANK2
|
ankyrin 2, neuronal
|
|
chr2_+_223625105
|
0.240
|
NM_080671
|
KCNE4
|
potassium voltage-gated channel, Isk-related family, member 4
|
|
chr9_-_72673753
|
0.240
|
NM_001007470
NM_020952
NM_024971
NM_206944
NM_206945
NM_206946
NM_206947
NM_206948
|
TRPM3
|
transient receptor potential cation channel, subfamily M, member 3
|
|
chr9_-_113561613
|
0.238
|
NM_001080551
|
C9orf84
|
chromosome 9 open reading frame 84
|
|
chr4_-_47831002
|
0.236
|
NM_003328
|
TXK
|
TXK tyrosine kinase
|
|
chr13_-_99422163
|
0.235
|
NM_033132
|
ZIC5
|
Zic family member 5 (odd-paired homolog, Drosophila)
|
|
chr5_-_20024050
|
0.235
|
NM_001167667
NM_004934
|
CDH18
|
cadherin 18, type 2
|
|
chr3_-_113495763
|
0.235
|
NM_183061
|
SLC9A10
|
solute carrier family 9, member 10
|
|
chr4_+_39084867
|
0.235
|
NM_175737
|
KLB
|
klotho beta
|
|
chr12_+_15017222
|
0.234
|
NM_006205
|
PDE6H
|
phosphodiesterase 6H, cGMP-specific, cone, gamma
|
|
chr8_+_26206648
|
0.234
|
NM_001177591
|
PPP2R2A
|
protein phosphatase 2, regulatory subunit B, alpha
|
|
chr6_-_128263868
|
0.234
|
NM_001010923
NM_001164685
|
THEMIS
|
thymocyte selection associated
|
|
chr16_-_21078262
|
0.230
|
NM_017539
|
DNAH3
|
dynein, axonemal, heavy chain 3
|
|
chr1_+_150937463
|
0.228
|
NM_178428
|
LCE2A
|
late cornified envelope 2A
|
|
chr2_-_213723298
|
0.227
|
NM_016260
|
IKZF2
|
IKAROS family zinc finger 2 (Helios)
|
|
chr1_+_84539876
|
0.226
|
NM_001134663
|
SAMD13
|
sterile alpha motif domain containing 13
|
|
chr3_+_112934016
|
0.226
|
NM_001134437
|
PHLDB2
|
pleckstrin homology-like domain, family B, member 2
|
|
chr4_+_46728051
|
0.224
|
NM_000812
|
GABRB1
|
gamma-aminobutyric acid (GABA) A receptor, beta 1
|
|
chr12_-_119038995
|
0.223
|
NM_001167606
NM_006861
|
RAB35
|
RAB35, member RAS oncogene family
|
|
chr12_-_69317441
|
0.222
|
NM_001109754
|
PTPRB
|
protein tyrosine phosphatase, receptor type, B
|
|
chr1_+_200698493
|
0.221
|
NM_032103
NM_032104
|
PPP1R12B
|
protein phosphatase 1, regulatory (inhibitor) subunit 12B
|
|
chr12_-_8584658
|
0.221
|
NM_014358
|
CLEC4E
|
C-type lectin domain family 4, member E
|
|
chr1_-_156056456
|
0.219
|
NM_001159397
NM_001159398
NM_052938
|
FCRL1
|
Fc receptor-like 1
|
|
chr10_+_50177192
|
0.218
|
NM_001135196
NM_199459
|
C10orf71
|
chromosome 10 open reading frame 71
|
|
chr3_+_188568861
|
0.217
|
NM_022147
|
RTP4
|
receptor (chemosensory) transporter protein 4
|
|
chr3_-_152449591
|
0.214
|
NM_001081455
|
P2RY14
|
purinergic receptor P2Y, G-protein coupled, 14
|
|
chr2_-_40593074
|
0.213
|
NM_001112802
|
SLC8A1
|
solute carrier family 8 (sodium/calcium exchanger), member 1
|
|
chr18_+_19706979
|
0.213
|
NM_000227
NM_001127718
|
LAMA3
|
laminin, alpha 3
|
|
chr20_+_44180296
|
0.213
|
NM_001250
NM_152854
|
CD40
|
CD40 molecule, TNF receptor superfamily member 5
|
|
chr10_-_5436781
|
0.213
|
NM_001171864
NM_024803
|
TUBAL3
|
tubulin, alpha-like 3
|
|
chr10_-_123724251
|
0.212
|
|
NSMCE4A
|
non-SMC element 4 homolog A (S. cerevisiae)
|
|
chr11_-_4902144
|
0.211
|
NM_001005237
|
OR51G1
|
olfactory receptor, family 51, subfamily G, member 1
|
|
chr2_+_228045053
|
0.209
|
|
AGFG1
|
ArfGAP with FG repeats 1
|
|
chr5_-_35974493
|
0.209
|
NM_001042625
NM_144647
|
CAPSL
|
calcyphosine-like
|
|
chr5_+_70787371
|
0.208
|
|
BDP1
|
B double prime 1, subunit of RNA polymerase III transcription initiation factor IIIB
|
|
chr15_+_72687765
|
0.207
|
NM_003992
|
CLK3
|
CDC-like kinase 3
|
|
chr2_+_219432789
|
0.206
|
NM_006522
|
WNT6
|
wingless-type MMTV integration site family, member 6
|
|
chr6_+_32021899
|
0.206
|
|
CFB
|
complement factor B
|
|
chr1_-_115039614
|
0.205
|
NM_000036
NM_001172626
|
AMPD1
|
adenosine monophosphate deaminase 1
|
|
chr9_-_21192201
|
0.205
|
NM_021057
|
IFNA7
|
interferon, alpha 7
|
|
chr2_-_36678657
|
0.204
|
NM_001042548
NM_005102
|
FEZ2
|
fasciculation and elongation protein zeta 2 (zygin II)
|
|
chr4_+_148758014
|
0.204
|
|
TMEM184C
|
transmembrane protein 184C
|
|
chr2_-_183095308
|
0.204
|
NM_005019
NM_001003683
|
PDE1A
|
phosphodiesterase 1A, calmodulin-dependent
|
|
chr9_-_21340885
|
0.204
|
NM_021002
|
IFNA6
|
interferon, alpha 6
|
|
chr1_+_48460943
|
0.203
|
NM_001011547
NM_001135181
|
SLC5A9
|
solute carrier family 5 (sodium/glucose cotransporter), member 9
|
|
chr11_-_64327197
|
0.201
|
NM_004579
|
MAP4K2
|
mitogen-activated protein kinase kinase kinase kinase 2
|
|
chr18_-_55136673
|
0.200
|
NM_181654
|
CPLX4
|
complexin 4
|
|
chr14_+_75058518
|
0.200
|
NM_006399
|
BATF
|
basic leucine zipper transcription factor, ATF-like
|
|
chr1_-_150328153
|
0.200
|
NM_001008536
|
TCHHL1
|
trichohyalin-like 1
|
|
chr2_+_176761552
|
0.199
|
NM_024501
|
HOXD1
|
homeobox D1
|
|
chr5_+_147628567
|
0.199
|
NM_001040129
|
SPINK13
|
serine peptidase inhibitor, Kazal type 13 (putative)
|
|
chr3_-_123765767
|
0.199
|
NM_001146102
NM_001146105
NM_001146104
NM_001146106
|
PARP9
|
poly (ADP-ribose) polymerase family, member 9
|
|
chr14_-_99911409
|
0.198
|
|
WARS
|
tryptophanyl-tRNA synthetase
|
|
chr3_-_197422653
|
0.197
|
NM_001039617
|
ZDHHC19
|
zinc finger, DHHC-type containing 19
|
|
chr1_-_150653351
|
0.196
|
NM_016190
|
CRNN
|
cornulin
|
|
chr11_+_109468734
|
0.196
|
|
ZC3H12C
|
zinc finger CCCH-type containing 12C
|
|
chr5_+_59819761
|
0.195
|
|
PART1
|
prostate androgen-regulated transcript 1 (non-protein coding)
|
|
chr2_+_169637255
|
0.195
|
NM_001142270
|
DHRS9
|
dehydrogenase/reductase (SDR family) member 9
|
|
chr17_-_10265991
|
0.193
|
NM_002472
|
MYH8
|
myosin, heavy chain 8, skeletal muscle, perinatal
|
|
chr1_-_142559339
|
0.192
|
NM_001123068
|
PPIAL4G
|
peptidylprolyl isomerase A (cyclophilin A)-like 4G
|
|
chr2_-_171796069
|
0.191
|
NM_001136554
|
TLK1
|
tousled-like kinase 1
|
|
chr15_+_70193652
|
0.189
|
NM_001166340
|
SENP8
|
SUMO/sentrin specific peptidase family member 8
|
|
chr5_-_41107161
|
0.189
|
NM_173489
|
HEATR7B2
|
HEAT repeat family member 7B2
|
|
chr10_+_50487146
|
0.189
|
NM_020984
|
CHAT
|
choline O-acetyltransferase
|
|
chr1_-_120155581
|
0.188
|
NM_001159352
NM_001159353
NM_032044
|
REG4
|
regenerating islet-derived family, member 4
|
|
chr5_+_66160359
|
0.188
|
NM_015183
|
MAST4
|
microtubule associated serine/threonine kinase family member 4
|
|
chr1_+_164063276
|
0.186
|
NM_012474
|
UCK2
|
uridine-cytidine kinase 2
|
|
chr4_-_122304925
|
0.186
|
NM_024873
|
TNIP3
|
TNFAIP3 interacting protein 3
|
|
chr14_+_75058591
|
0.185
|
|
BATF
|
basic leucine zipper transcription factor, ATF-like
|
|
chr14_-_72563497
|
0.185
|
NM_021260
|
ZFYVE1
|
zinc finger, FYVE domain containing 1
|
|
chr2_+_218641982
|
0.182
|
NM_198483
|
RUFY4
|
RUN and FYVE domain containing 4
|
|
chr2_-_39310176
|
0.181
|
NM_001009565
|
CDKL4
|
cyclin-dependent kinase-like 4
|
|
chr3_-_152173475
|
0.180
|
NM_001195794
NM_174878
|
CLRN1
|
clarin 1
|
|
chr3_+_38238080
|
0.180
|
|
OXSR1
|
oxidative-stress responsive 1
|
|
chr2_+_234624084
|
0.179
|
NM_006944
|
SPP2
|
secreted phosphoprotein 2, 24kDa
|
|
chr2_-_134042500
|
0.178
|
NM_207363
NM_207481
|
NCKAP5
|
NCK-associated protein 5
|
|
chr2_-_216944911
|
0.177
|
NM_020814
|
MARCH4
|
membrane-associated ring finger (C3HC4) 4
|
|
chr7_-_150555153
|
0.176
|
NM_005692
NM_007189
|
ABCF2
|
ATP-binding cassette, sub-family F (GCN20), member 2
|
|
chr3_+_63928459
|
0.176
|
NM_001128149
|
ATXN7
|
ataxin 7
|
|
chr17_-_1195096
|
0.175
|
|
PAFAH1B1
|
platelet-activating factor acetylhydrolase 1b, regulatory subunit 1 (45kDa)
|
|
chr12_-_119038940
|
0.175
|
|
RAB35
|
RAB35, member RAS oncogene family
|
|
chr4_-_39199527
|
0.175
|
|
UGDH
|
UDP-glucose 6-dehydrogenase
|
|
chr3_-_9266251
|
0.174
|
NM_001033117
NM_014850
|
SRGAP3
|
SLIT-ROBO Rho GTPase activating protein 3
|
|
chr5_+_6799100
|
0.172
|
NM_001171806
|
PAPD7
|
PAP associated domain containing 7
|
|
chr1_+_40977092
|
0.171
|
|
NFYC
|
nuclear transcription factor Y, gamma
|
|
chr2_-_97978421
|
0.170
|
NM_015348
|
TMEM131
|
transmembrane protein 131
|
|
chr12_-_2856356
|
0.170
|
|
FOXM1
|
forkhead box M1
|
|
chr20_+_44180355
|
0.170
|
|
CD40
|
CD40 molecule, TNF receptor superfamily member 5
|
|
chr14_-_106184988
|
0.169
|
|
IGHA1
IGHG1
|
immunoglobulin heavy constant alpha 1
immunoglobulin heavy constant gamma 1 (G1m marker)
|
|
chr8_+_120148604
|
0.169
|
NM_006438
|
COLEC10
|
collectin sub-family member 10 (C-type lectin)
|
|
chr12_+_5023345
|
0.169
|
NM_002234
|
KCNA5
|
potassium voltage-gated channel, shaker-related subfamily, member 5
|
|
chr6_+_31647854
|
0.169
|
NM_001159740
|
LTA
|
lymphotoxin alpha (TNF superfamily, member 1)
|
|
chr22_-_25205376
|
0.168
|
NM_152841
|
HPS4
|
Hermansky-Pudlak syndrome 4
|
|
chr7_-_86433139
|
0.166
|
NM_152748
|
KIAA1324L
|
KIAA1324-like
|
|
chr12_+_119039075
|
0.166
|
|
|
|
|
chr3_+_35697432
|
0.164
|
NM_198399
|
ARPP21
|
cAMP-regulated phosphoprotein, 21kDa
|
|
chr4_-_40211324
|
0.164
|
|
RBM47
|
RNA binding motif protein 47
|
|
chr3_-_69381995
|
0.164
|
|
FRMD4B
|
FERM domain containing 4B
|